KEYWORDS
Microbiome
Gut microbiome
in vitro
in vivo
in silico
health
Metabolic
Joint
Immunity
Abstract
A healthy diet has been recognized as essential to human nutrition and health, but there is still much debate as to what entails a healthy diet. A large part of the health effect of a diet is related to the metabolization capacity of the gut microbiome, as metabolites produced by the gut bacteria can affect the host. Many of the interactions between diet, gut microbiome and host that influence human health are still unknown, in part due to a lack in mechanistic insights into the metabolization potential of the gut microbiome. This metabolization potential can be studied in more detail in in vitro systems combined with metagenomic in silico analysis. This combined strategy provides mechanistic insights and leads for diet intervention studies to verify if these mechanisms also occur in vivo. Another, more novel, in silico-approach using artificial intelligence models can also help to understand the complex mechanisms of the gut microbiome and its influence on human health. All these strategies combined will help define what constitutes a healthy, plant-forward diet.

Diet and dietary components can direct the gut microbiome composition and functionality
A healthy diet has been recognized as essential to human nutrition and health (1). Despite the extensive research on the influence of diet on human health, there are still many controversies on what entails a healthy diet. This is in part due to the limited consideration of the interactions between diet, gut microbiome and host (2). The habitual diet determines, among other factors such as medication, age, or geography, the gut microbial community as well as its metabolic capacity (3–5). The gut microbiome has an influence on what the human host is able to extract from the diet and the metabolization potential of the gut microbiome can impact human health by contributing to host metabolism and immunology (6). The selective nature of dietary components can be used to direct the gut microbiome towards the production of health-promoting metabolites (2,7,8). To do so, we need a better understanding of microbial community dynamics and underlying selective mechanisms, e.g. interspecies competition, in order to optimally use dietary components for the stimulation of health promoting metabolites (9,10).
The effect of a plant-forward diet on the gut microbiome
Understanding the impact of diet on the gut microbiome is not only important from a human health perspective, but it is also important in light of planetary health and the necessary transfer towards a more plant-forward diet (11). It can be expected that an increasingly plant-based diet will contain more fiber that could benefit the gut microbiome, and more knowledge on this will help with the development of sustainable gut-friendly plant-based products required for this transfer (2,12). Additionally, when people get more insights in the beneficial effect of a plant-forward diet on their own gut microbiome, it can motivate them to make long-lasting dietary changes towards healthier and more sustainable diets (13).
One of the aspects of a more plant-forward diet is that plant material contains more fibers, both digestible as well as non-digestible, compared to animal-based material (8,14). The majority of non-digestible fibers pass through the digestion system unaltered, until they reach the colon where the gut microbial community can use them as substrate. The community shaping properties of non-digestible fibers has been recognized, but there is still a knowledge-gap of how the structure of fibers shape the function of the gut microbial community, which results in a low predictability (14). Interestingly, it has been shown that the change in metabolic functionality was relatively predictable despite the highly individual change in gut microbiome composition when presented with a fiber-rich diet (15). This suggests that a functional-focussed approach, instead of a species-focussed approach, could decrease the level of complexity of the interplay between diet, health and gut microbiome and increase our understanding. Additionally, this will help to create more mechanistic insights into the selectiveness of dietary components and potentially help to provide more personalized dietary advice that benefits people's gut microbiome and thereby their health.

In vitro model systems as a screening tool to investigate the effect of substrates on the gut microbiome
The preferred way of obtaining mechanistic understanding of the gut microbiome modulating potential is via human dietary intervention trials. However, these types of studies are relatively expensive, especially when used to test many products and combined with a multi-omics microbiome approach. Animal models can also be used to investigate the effect of diet on health, but direct translation to the human situation is difficult because of different genetics and microbiome community composition (16,17). Alternatives to animal models are in vitro model systems to investigate specific substrate-microbiome relationships. There are multiple options for these types of model systems, with different levels of complexity. The simplest model system, a batch (fermentation) system, allows for rapid high-throughput screening of short-term metabolization of substrates. However, the lack of a representative environment, e.g. no mucus layer nor host absorption, reduces the direct translatability to the human gut. Other more complex intestinal models which mimic the digestive system, The TNO intestinal model (TIM-2) or the Simulated Human Intestinal Microbial Ecosystem (SHIME), are more representative for the dynamic gut microbiome environment (16,18). However, these models are more limited in the number of conditions which can be tested in a short timeframe compared to the batch systems. No matter which system is used, ultimately, results should be validated in human in vivo intervention trials, to check e.g. whether dose is properly chosen, tolerability is not an issue, and whether the (predicted) health effect can be confirmed.
Using in silico-based strategies to determine the gut microbiome capacity to process nutrients, foods and diet
So far, many gut microbiome in vitro studies have predominantly focused on microbial community composition shifts on genus-level and the correlation with the production of simple, health-relevant metabolites (e.g. short-chain fatty acids, p-cresol, branch-chain amino acids). Unfortunately, these types of studies often lack a mechanistic insight how, via which microbial metabolic pathway, these metabolites were produced. As many of the health effects are elicited by the metabolites produced by the gut microbiome, mechanistic insights are essential to understand the effect of food on human health (17,19). A way to gain more insight into the metabolic potential of the microbial community is to combine an insilico-based metagenomic analysis with the above-mentioned studies. Due to their sampling accessibility, in vitro systems, especially compared to in vivo, are ideal for these types of combined approaches (18). An in silico approach would be to map a specific metabolization potential of the microbiome of interest before and after incubation with specific substrates. By investigating the differential count of genes of interest it is possible to identify which of the functionalities, and by default species, thrived under the imposed conditions (Figure 1). Another in silico strategy would be to mine the publicly available gut microbiome databases for a specific metabolization potential of interest. This can result in a list of potentially produced metabolites or commensal species which could have a beneficial health effect for the host (Figure 1). This information can then be used to support gut-friendly products development or design a targeted in vivo trial to verify the potential health benefits.
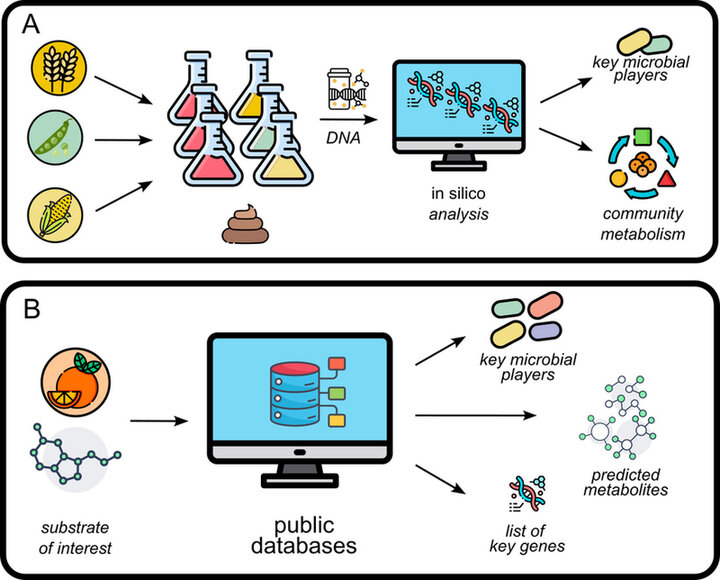
Figure 1 | A. In silico strategy to identify key species and community metabolism of specific substrates (e.g. a specific fiber) tested in in vitro systems inoculated with a fecal community. B.In silico strategy to investigate a substrate or metabolization potential of interest and identify associated commensal species, metabolites and key genes. Icons derived from: https://www.flaticon.com/.
Next steps in in silico modelling; using artificial intelligence to predict the cazyme response of the gut microbiome to specific dietary fibers
The microbial degradation potential of dietary fibers has been extensively studied, and although there are still a lot of knowledge-gaps in the structure-function relationship between fibers and the gut community, also a lot is already known. This available functional knowledge is combined in a large database, the carbohydrate active enzyme (CAZyme) database, which is publicly available to explore and extend our insights into structure-function relationships (20). These CAZymes are employed by bacteria to degrade fibers which enter the colon and produce bioactive metabolites in the process (21,22). Indeed, studies have shown that personal differences in fiber-based diet interventions are driven by the gut microbiome CAZyme repertoire (15,22). A new project, FIBERME (LWV23141), has started at Wageningen Food & Biobased Research, and aims to develop an artificial intelligence (AI)-model, which, based on the substrate-CAZyme relationship, predicts the microbiome response to fiber intake. Alternatively, it can predict the shift in microbiome composition based on a given supplied fibre (Figure 1). This ambitious project will perform several thousands of incubations, supplemented with already existing data to understand the mechanistic effect of dietary fibers on the gut microbiome from a point of view of the structure of the fibre (which is linked to the induced CAZyme profile). It will generate targeted in vitro and in vivo data to increase the accuracy of predictions. The final model will be turned into an interactive AI-tool. Both the envisioned AI-tool and gained knowledge will provide guidance to food industry in the development of gut-friendly foods and potentially support accurate personalized dietary advice on fiber intake and accompanying health benefits. For instance, the tool can help in the selection of the best fiber for an optimal health effect. In this way FIBERME can contribute to a more targeted and health supporting increase in fiber intake in the population. Indirectly, it will also facilitate the plant-protein transition and limit food waste by understanding how to transform fiber-rich plant-based waste streams into highly valuable products.

Conclusions
Although it has been well established that diet has an impact on human health, there is still much discussion about what constitutes a healthy diet. This is mostly due to the limited understanding of the interplay between the diet, gut microbiome and human health. Getting a more mechanistic insight into the effect of diet on the gut microbiome and particularly the metabolites produced by the gut microbiome will help to develop more healthy diets. This understanding is also important in light of the required shift towards a more plant-based diet. Although in vivo studies are the ultimate system to test the health effect of diets, due to the costs of those studies other methods are needed to pre-select dietary components with a proposed health effect. Using an approach which combines genome-based in silico analysis with in vitro systems will allow for high-throughput testing of substrate-function relationships of the gut microbial community. Additionally, this approach allows for mechanistic understanding of how the substrates were converted to the health-affecting metabolites. A step further is to develop an AI-model which predicts the gut microbial community response to specific fibers. All these steps combined with help define what really constitutes a healthy plant-forward diet.
Gut-skin-axis
We will see amla listed if we refer to the list of ingredients with specific effects on gut health. Amla is also high in polyphenols, and research shows it is an excellent skin health, primarily in control of collagen metabolism. (22)
Conclusion
The growing body of evidence showing that botanicals play an important role directly on gut health and the microbiome is promising. With new research stating the impact of botanicals high in polyphenols on gut health and the microbiome leads us to ask the question, why not combine botanicals with probiotics to achieve a better outcome for the gut, and gut-blank-axis health concerns? Based on the growing body of evidence, botanicals should be considered a primary way to support both gut and gut-blank-axis health concerns.
References and notes
- Jaacks LM, Vandevijvere S, Pan A, McGowan CJ, Wallace C, Imamura F, et al. The obesity transition: stages of the global epidemic. The Lancet Diabetes & Endocrinology. 2019 Mar;7(3):231–40.
- Armet AM, Deehan EC, O’Sullivan AF, Mota JF, Field CJ, Prado CM, et al. Rethinking healthy eating in light of the gut microbiome. Cell Host & Microbe. 2022 Jun;30(6):764–85.
- Yatsunenko T, Rey FE, Manary MJ, Trehan I, Dominguez-Bello MG, Contreras M, et al. Human gut microbiome viewed across age and geography. Nature. 2012 Jun;486(7402):222–7.
- Rothschild D, Weissbrod O, Barkan E, Kurilshikov A, Korem T, Zeevi D, et al. Environment dominates over host genetics in shaping human gut microbiota. Nature. 2018 Mar;555(7695):210–5.
- Gacesa R, Kurilshikov A, Vich Vila A, Sinha T, Klaassen MAY, Bolte LA, et al. Environmental factors shaping the gut microbiome in a Dutch population. Nature. 2022 Apr 28;604(7907):732–9.
- de Vos WM, Tilg H, Van Hul M, Cani PD. Gut microbiome and health: mechanistic insights. Gut. 2022 May;71(5):1020–32.
- De Filippo C, Cavalieri D, Di Paola M, Ramazzotti M, Poullet JB, Massart S, et al. Impact of diet in shaping gut microbiota revealed by a comparative study in children from Europe and rural Africa. Proc Natl Acad Sci USA. 2010 Aug 17;107(33):14691–6.
- David LA, Maurice CF, Carmody RN, Gootenberg DB, Button JE, Wolfe BE, et al. Diet rapidly and reproducibly alters the human gut microbiome. Nature. 2014 Jan;505(7484):559–63.
- Wu GD, Compher C, Chen EZ, Smith SA, Shah RD, Bittinger K, et al. Comparative metabolomics in vegans and omnivores reveal constraints on diet-dependent gut microbiota metabolite production. Gut. 2016 Jan;65(1):63–72.
- Patnode ML, Beller ZW, Han ND, Cheng J, Peters SL, Terrapon N, et al. Interspecies competition impacts targeted manipulation of human gut bacteria by fiber-derived glycans. Cell. 2019 Sep;179(1):59-73.e13.
- Xu X, Sharma P, Shu S, Lin TS, Ciais P, Tubiello FN, et al. Global greenhouse gas emissions from animal-based foods are twice those of plant-based foods. Nat Food. 2021 Sep 13;2(9):724–32.
- Willett W, Rockström J, Loken B, Springmann M, Lang T, Vermeulen S, et al. Food in the Anthropocene: the EAT–Lancet Commission on healthy diets from sustainable food systems. The Lancet. 2019 Feb;393(10170):447–92.
- World Health Organization. Shanghai declaration on promoting health in the 2030 Agenda for Sustainable Development. Health Promotion International. 2017 Feb;32(1):7–8.
- Rastall RA, Diez-Municio M, Forssten SD, Hamaker B, Meynier A, Moreno FJ, et al. Structure and function of non-digestible carbohydrates in the gut microbiome. Beneficial Microbes. 2022 Jun 18;13(2):95–168.
- Delannoy-Bruno O, Desai C, Raman AS, Chen RY, Hibberd MC, Cheng J, et al. Evaluating microbiome-directed fibre snacks in gnotobiotic mice and humans. Nature. 2021 Jul 1;595(7865):91–5.
- Wilmes P, Calatayud M, Wiele TV de. Resolving host–microbe interactions in the gut: the promise of in vitro models to complement in vivo research. Current Opinion in Microbiology. 2018 Aug;44:28–33.
- Walter J, Armet AM, Finlay BB, Shanahan F. Establishing or exaggerating causality for the gut microbiome: lessons from human microbiota-associated rodents. Cell. 2020 Jan;180(2):221–32.
- Isenring J, Bircher L, Geirnaert A, Lacroix C. In vitro human gut microbiota fermentation models: opportunities, challenges, and pitfalls. Microbiome Res Rep. 2023;2(1):2.
- Makki K, Deehan EC, Walter J, Bäckhed F. The impact of dietary fiber on gut microbiota in host health and disease. Cell Host & Microbe. 2018 Jun;23(6):705–15.
- Drula E, Garron ML, Dogan S, Lombard V, Henrissat B, Terrapon N. The carbohydrate-active enzyme database: functions and literature. Nucleic Acids Research. 2022 Jan 7;50(D1):D571–7.
- Lapébie P, Lombard V, Drula E, Terrapon N, Henrissat B. Bacteroidetes use thousands of enzyme combinations to break down glycans. Nat Commun. 2019 May 3;10(1):2043.
- Kok CR, Rose D, Hutkins R. Predicting personalized responses to dietary fiber interventions: opportunities for modulation of the gut microbiome to improve health. Annu Rev Food Sci Technol. 2023 Mar 27;14(1):157–82.



